Author: Manuel J. Arroyo Pulgar
At Clover Biosoft, we develop methods to identify and classify resistant microorganisms reducing time to result and cost of other methods and reagents. But our technology can go much beyond that.
One of the advantages of mass spectrometry technology is its versatility, which can be used in many research fields, including cancer research. In fact, Clover Biosoft worked together with several departments in Medical University of Vienna (Austria) and Shimadzu Corporation (UK). The result of that work was the publication of the article MALDI-MS Protein Profiling of Chemoresistance in Extracellular Vesicles of Cancer Cells (Figure 1), one of the main works of Dr. Gerald Stübiger for which he was awarded researcher of the month at Medical University of Vienna [1].
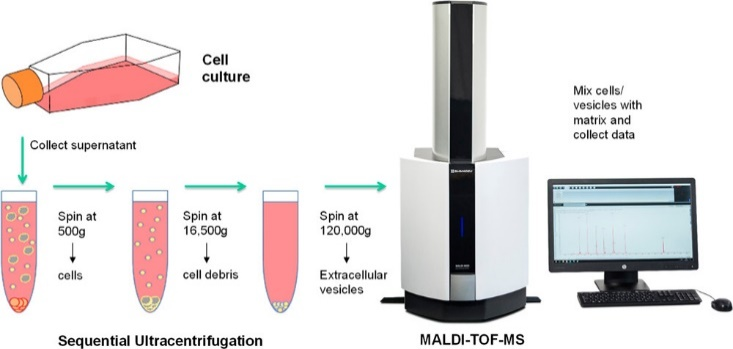
In this research [2] MALDI-TOF MS was used for protein profiling of extracellular vesicles (EVs) derived from a cellular model of chemoresistance in cancer cells comprising the CCL-228 cell line as the primary colon tumour, the lymph node metastasis CCL-227, and subclones resistant to different 5-fluorouracil concentrations. These EVs represent cellular organelles in the nanometre range derived from the plasma membrane or the endosome. A specific type of EV is exosomes representing small vesicles (30-120 nm) that are known to contain DNA, mRNA, miRNA, proteins lipids and viruses. Exosomes are found in raising concentrations in cancer patients where they can reprogram other cells in the organism to support malignant progression of the tumour. Thus, they represent very promising sources of novel biomarkers for non-invasive diagnostics and vectors of therapy, since all cells can produce them.
As the experiments showed, the quantities of EVs isolated from cell supernatants would be enough to establish a characteristic protein profile from a few millilitres of blood, using MALDI-MS. Our study opens the door to a completely new field of application for MALDI-MS in clinical diagnostics and beyond, explains Gerald Stübiger [3].
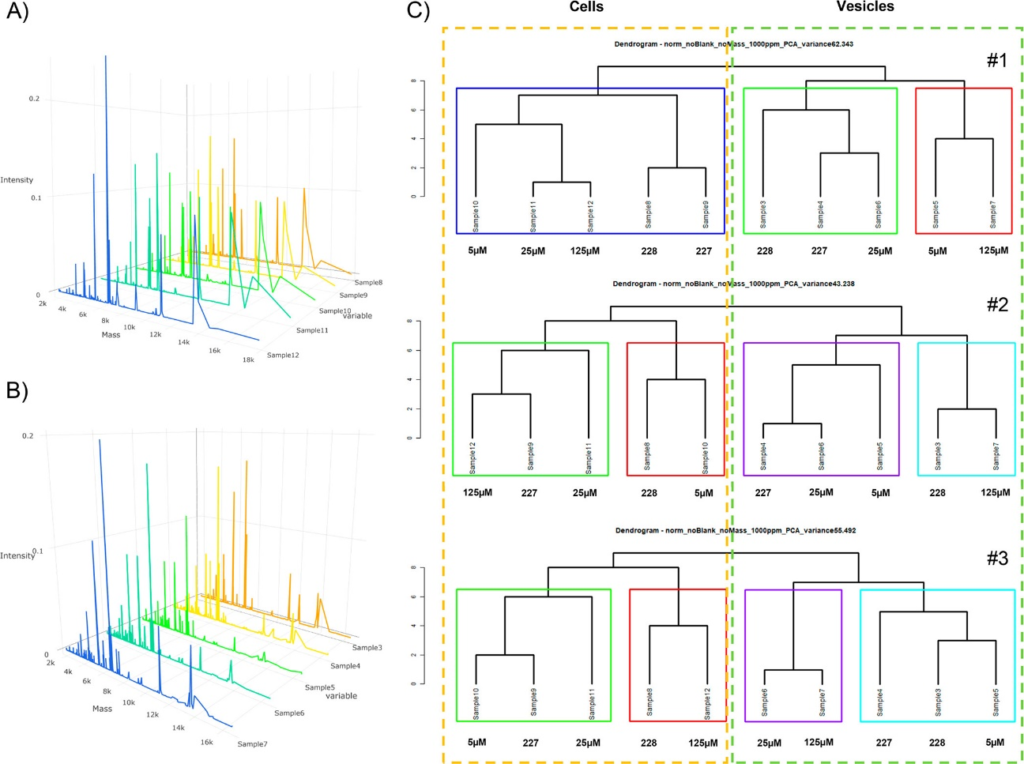
This work demonstrates the proof-of-concept that MALDI-MS protein profiling can be used to characterize EVs (e.g., exosomes) from different cancer cell populations and also between those expressing different levels of susceptibility to chemotherapy. This method is promising for application and validation in clinical settings. It is anticipated that applying this technique to minimally invasive diagnostics from body fluids (e.g., human plasma) will essentially contribute to improve cancer therapy monitoring in the future.
SOURCES:
[2] MALDI-MS Protein Profiling of Chemoresistance in Extracellular Vesicles of Cancer Cells | https://pubs.acs.org/doi/abs/10.1021/acs.analchem.8b03756
